Projects
Structure and mechanism of erythromycin resistance methyltransferases
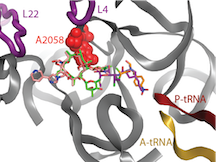
The active site of the large ribosomal subunit, the peptidyl transferase center (PTC), and its surrounding environment are important targets for antibiotic action. Enzymes, such as erythromycin resistance methyltransferase and chloramphenicol resistance methyltransferase, methylate specific residues adjacent to the PTC to block antibiotic binding and initiate resistance to multiple classes of antibiotics. Despite the medical importance of these enzymes, the mechanisms governing how they specifically recognize their target nucleotide and promote catalysis are not fully known. We aim to understand these mechanisms to aid in developing inhibitors to these resistance enzymes, restoring the functionality of existing antibiotic therapies.
Mechanisms and regulation of CRISPR-Cas interference |
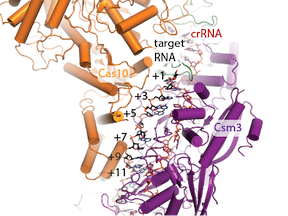
CRISPR-Cas is an RNA-protein complex found in many bacteria and archaea that defends the microbe against viruses and other foreign nucleic acids. Spacer genes (spc) are composed of snippets of sequence from a bacterial virus or foreign nucleic acid that attacked an ancestral cell. Transcription of spacer genes results in crRNA, a short RNA bound by Cas proteins that forms a functional complex able to detect and interfere with foreign nucleic acids complementary to the crRNA. Diverse CRISPR-Cas systems have been discovered that can be grouped into two classes and at least six types. We are studying the architecture, regulation and function of Type III CRISPR-Cas. We are interested in how CRISPR-Cas contributes to bacterial fitness and the engineering of CRISPR-Cas for biotechnology.
Structural biology of sulfur mobilization
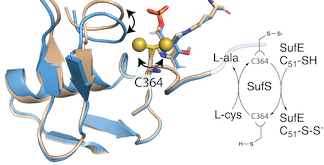
The amino acid cysteine serves as the major source for the entry of sulfur into biosynthesis pathways. Cysteine desulfurases are responsible for liberating sulfur from cysteine so that it can be used in the synthesis of sulfur-containing cofactors. We have performed x-ray crystallographic studies of the cysteine desulfurase, SufS, to understand its mechanism of activity and regulation. Our studies of cysteine desulfurase, an enzyme family present in all life, are clarifying how sulfur enters diverse biochemical pathways of the cell.

