|
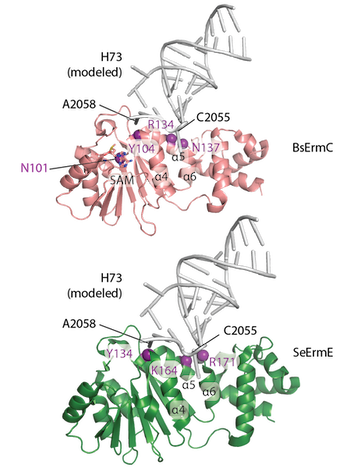
Erythromycin resistance methyltransferase (Erm) enzymes are a clinically important mechanism of multi-drug antibiotic resistance rendering bacteria resistant to macrolide, lincosamide and ketolide antibiotics. Whether or not all Erm enzymes bind to their rRNA substrate in a similar manner is an outstanding question. A new publication from the Dunkle group titled "Shared requirements for key residues in the antibiotic resistance enzymes ErmC and ErmE suggest a common mode of RNA recognition" appeared in the December 18 edition of the Journal of Biological Chemistry. The article demonstrates that similar conserved residues in ErmC or ErmE are required for function by deploying in vivo and in vitro assays on a collection of site-directed mutants. Also a structural model for how Erm enzymes bind rRNA is presented.
Caroline Yuk, an undergraduate researcher in the Dunkle group and UA Neuroscience major, is one of the 2020 recipients of the Marshall Scholarship. Caroline was among the 46 U.S. students chosen in 2020 and will complete two years of graduate studies in the United Kingdom working toward her goal of becoming a medical researcher. Link to the UA press release.
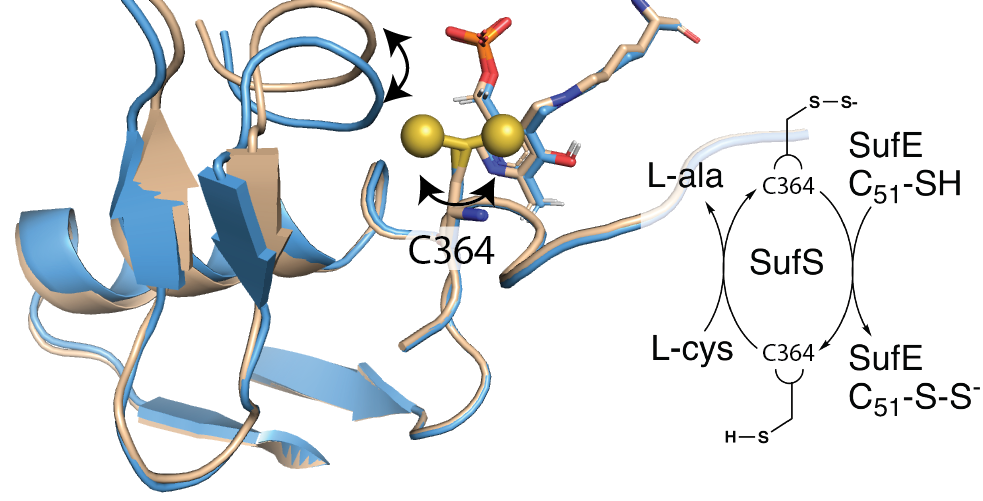
A recent research article from the Dunkle group 'Structural evidence for dimer-interface driven regulation of the type II cysteine desulfurase, SufS' was listed among the most cited mechanistic enzymology articles appearing in the journal Biochemistry (published by the American Chemical Society) in 2019. Here's the link. Research on the Suf pathway for iron-sulfur cluster biogenesis is performed in collaboration with the Frantom group at the University of Alabama and the Outten group at the University of South Carolina.
|
|
|
Accessibility | Equal Opportunity | UA Disclaimer | Site Disclaimer | Privacy | Copyright © 2020
The University of Alabama | Tuscaloosa, AL 35487 | (205) 348-6010 Website provided by the Center for Instructional Technology, Office of Information Technology |

 RSS Feed
RSS Feed
